|
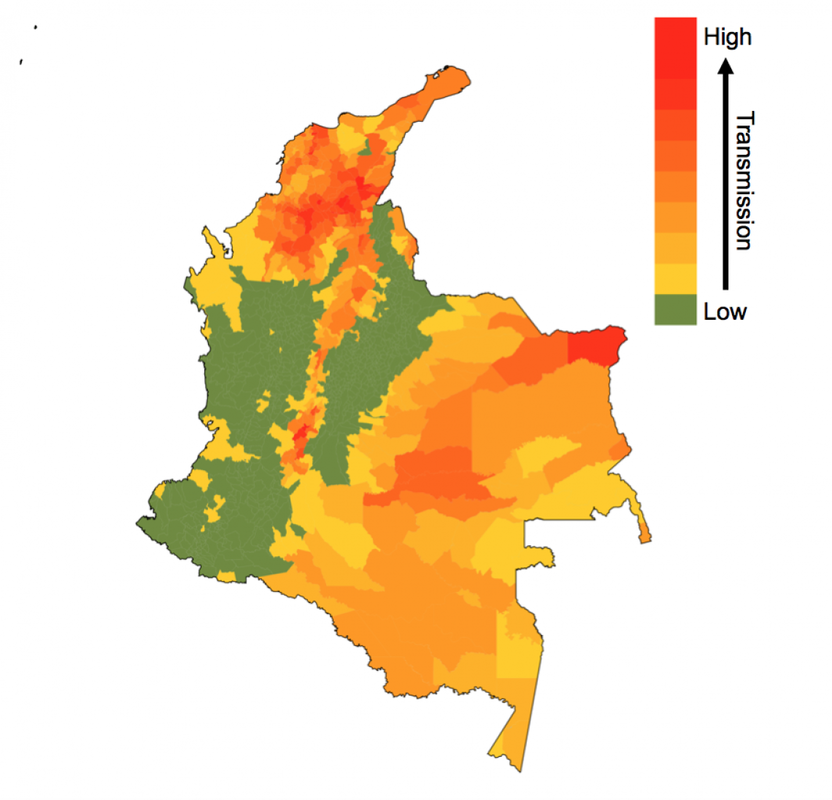
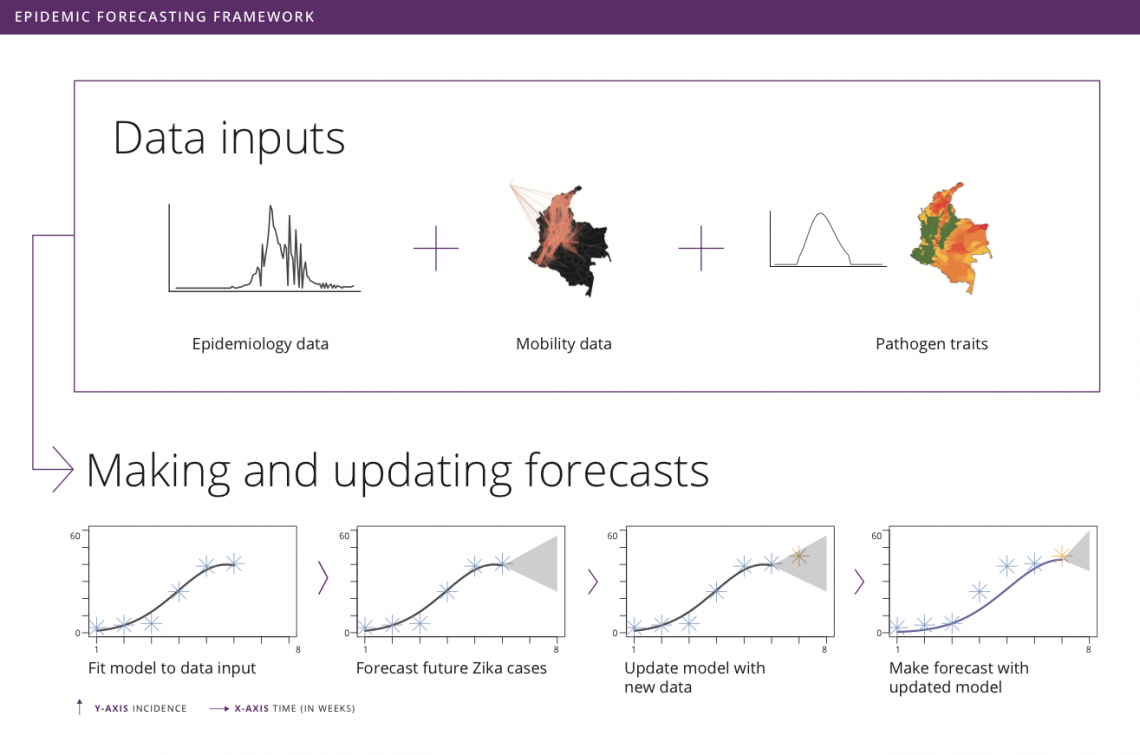
by Rachel Oidtman, Alex Perkins, Moritz Kraemer, Manuel Garcia-Herranz, and Elisa Omodei Originally published on: https://www.unicef.org/innovation/stories/epidemicforecasting The decision to grab your umbrella as you head off to work in the morning is shaped by the daily weather forecast. Although meteorology has been a formal science since the 19th century, it was only in the last 20 years that these forecasts became accurate enough (>60% accuracy) that we could trust them and use weather forecasts to make decisions. In the background of our daily lives, there has been a “quiet revolution” in weather forecasting, kick-started by a number of technological and scientific advances over the last few decades. With considerable technological advancements across all scientific disciplines, including a vast increase in the amount of data and more realistic models, meteorology is the only field that has achieved such clear success when it comes to forecasting. This begs the question: What if we could forecast influenza with the accuracy of rain forecasts? Or, what if we could forecast an emerging disease with the accuracy of hurricane forecasts? Weather forecasting as a model for epidemic forecasting With hurricanes, meteorologists know the seasons in which they occur, but they cannot begin forecasting the progression of a hurricane system until the first signs of a storm appear (i.e. low pressure system with thunderstorms during hurricane months). Early on in hurricane forecasts, there is significant uncertainty surrounding both the spatial trajectory and severity of the storm. With weekly weather forecasts, meteorologists know seasonal weather patterns and have extensive geographic data on weather variables. There is uncertainty around weekly weather forecasts, but this is much more constrained than hurricane forecasts. Building and improving upon both hurricane and weekly weather forecasts, meteorologists collect more data, invest in better satellites (collecting more precise data), and consider more models. Learning from the ways in which meteorologists improved both extreme and seasonal weather forecasts, it becomes clear that better models and more data could help to improve disease forecasting. Traditional disease surveillance data though cannot be collected as efficiently and consistently as weather data. Hence, the scientific community is devising more creative options to improve disease surveillance and epidemic forecasting. One such option is to use unconventional data sources (e.g. human mobility data, social media traces, web searches, etc.) to represent biological processes (e.g. pathogen dispersion) to enhance the realism of assumptions about disease spread in forecasts. Another complementary option is to organize forecasting “challenges”, which promote the development of more accurate models and offer a means to compare the accuracies of different models. Forecasting familiar and foreign foes Even with the sparse and varied nature of epidemiological data, there has nonetheless been progress in disease forecasting of both recurring seasonal diseases and emerging diseases. “Familiar foes” that exhibit recurring, seasonal transmission, such as influenza and dengue, have received considerable attention. One of the first major initiatives was Google Flu Trends, launched in 2008, which used Google search queries to forecast flu incidence. Since then, there has been consistent progress as academic researchers have been steadfastly working on developing more and more accurate models. The most notable example is the CDC-developed flu forecasting initiative, “FluSight”, that is used as a platform for visualizing and sharing real-time weekly flu forecasts contributed by various modeling teams around the world. Dengue has also been at the center of attention of the scientific community since the Dengue Forecasting Project was launched in 2015 - a forecasting contest based on historical data. “Foreign foes” that appear with little warning and can spread rapidly across international borders remain a more elusive challenge for disease forecasting, with recent notable examples including Ebola and Zika viruses. Chikungunya, a mosquito-borne disease that spread across the Americas in 2013-2015, offers another example. While retrospective analyses indicated some success with forecasting the sequence in which different islands in the Caribbean would be invaded, a near real-time forecasting challenge sponsored by Defense Advanced Research Projects Agency (DARPA) showed that most models tended to have difficulty forecasting other features of the epidemic, such as the week in which incidence would peak. Forecasting was also performed in support of the Ebola epidemic around the same time in West Africa, although somewhat controversially. Human movement and forecasting A variety of biological characteristics among different pathogens may contribute to variation in how difficult they are to forecast. For example, outbreaks of some diseases depend almost exclusively on human contact (e.g. influenza), whereas others involve frequent spillover from animals (e.g. MERS). One way to advance the science of forecasting across such a wide range of infectious diseases is to focus on challenges that are common to forecasting of many pathogens. One such challenge is capturing the role of human movement in pathogen spread. Consider an emerging disease, such as Zika. Early on in its epidemic in the Americas, there were a very limited number of locations that had confirmed the presence of Zika virus. Theoretically, if there was no human movement in or out of these locations, then the pathogen would have been restricted to these isolated locations (mosquitoes are also involved in Zika virus transmission but tend not to move very far). Because there is regular movement between locations, Zika virus can spread to new locations with humans as they travel for work, leisure, etc. Nowadays, this is possible thanks to the huge amount of data that each of us generates using portable technology. Every time we make a phone call or post a picture on Instagram, we are recording our location. If today someone makes a call from Bogotá, and tomorrow a call from Cali, the phone company knows that one person has traveled from one city to the other. Aggregating this information across many users, telephone companies can provide UNICEF and other stakeholders with anonymized data on human mobility patterns across an entire country in real time. Forecasting vector-borne disease outbreaks at UNICEF Since spring 2018, the Office of Innovation at UNICEF has been working with academic researchers at University of Notre Dame and Boston Children’s Hospital to meld together epidemic modeling approaches with innovative data sets to help the most vulnerable. Together, we developed a forecasting model for Zika in Colombia’s more than one thousand municipalities. In each municipality, we use a basic model that takes into account the environmentally driven nature of Zika virus transmission, and the accumulation of population immunity as the epidemic grows, to generate incidence forecasts. Across municipalities, we aim at modelling the spread of Zika virus using human mobility patterns that can be obtained from anonymized and aggregated cell phone data like that provided by UNICEF’s partner, Telefónica. To explore the real-time capabilities of this forecasting model, we iteratively fit the model, make forecasts, and update our model as new incidence data becomes available. As this collaborative venture continues together with UNICEF Colombia, our goal is to develop a user-friendly interface where public health officials from the Colombian Ministry of Health and National Institute of Health can plug in surveillance data and use our technically sophisticated modeling machinery to make forecasts for future Zika epidemics. This information will be used to make faster decisions on planning and prioritizing preparedness and response actions, thereby helping prevent the spread of such epidemics. The plan is to soon extend this work to include forecasts for other vector-borne diseases currently affecting Colombia, such as dengue and malaria. These forecasting tools will become a core component of MagicBox - UNICEF’s open-source software platform that enables collaboration and the use of new data sources and computational techniques, like AI and machine learning, for good. Recent Zika epidemics in Angola and India serve as a reminder that the threat posed by Zika is, unfortunately, not over. In this regard, the results from this collaboration will have direct implications for policy and public health.
This project builds on prior research funded by a RAPID grant from the National Science Foundation and a Branco Weiss Fellowship. 8/1/2022 06:43:18 am
Really informative article, I had the opportunity to learn a lot, thank you. https://takipcisatinalz.com/takipci-2/ 8/2/2022 04:03:53 am
Really informative article, I had the opportunity to learn a lot, thank you. https://www.ugurelektronik.com/ 8/2/2022 09:47:03 am
Really informative article, I had the opportunity to learn a lot, thank you. https://takipcialdim.com/ucuz-takipci-satin-al/ Comments are closed.
|